Note
Go to the end to download the full example code.
1.1. YAML-Based Model Definitions
In this tutorial, you will learn how to use YAML definition files to implement a neurodynamic model with multiple interconnected neural populations. In this process, the following questions will be answered:
What is a YAML template file?
How do I use the YAML templates to define my model
What are the roles of the different types of YAML templates used in PyRates?
How do I create a network model from a YAML template in Python?
Throughout the tutorial, we will make use of the Jansen-Rit neural mass model [1], which describes the dynamic interactions between 3 neural populations via their average firing rates. It has been used to describe the macroscopic, electrophysiological activity within a cortical column. A graphic representation of such a model, placed inside a brain network, can be found in the figure below.
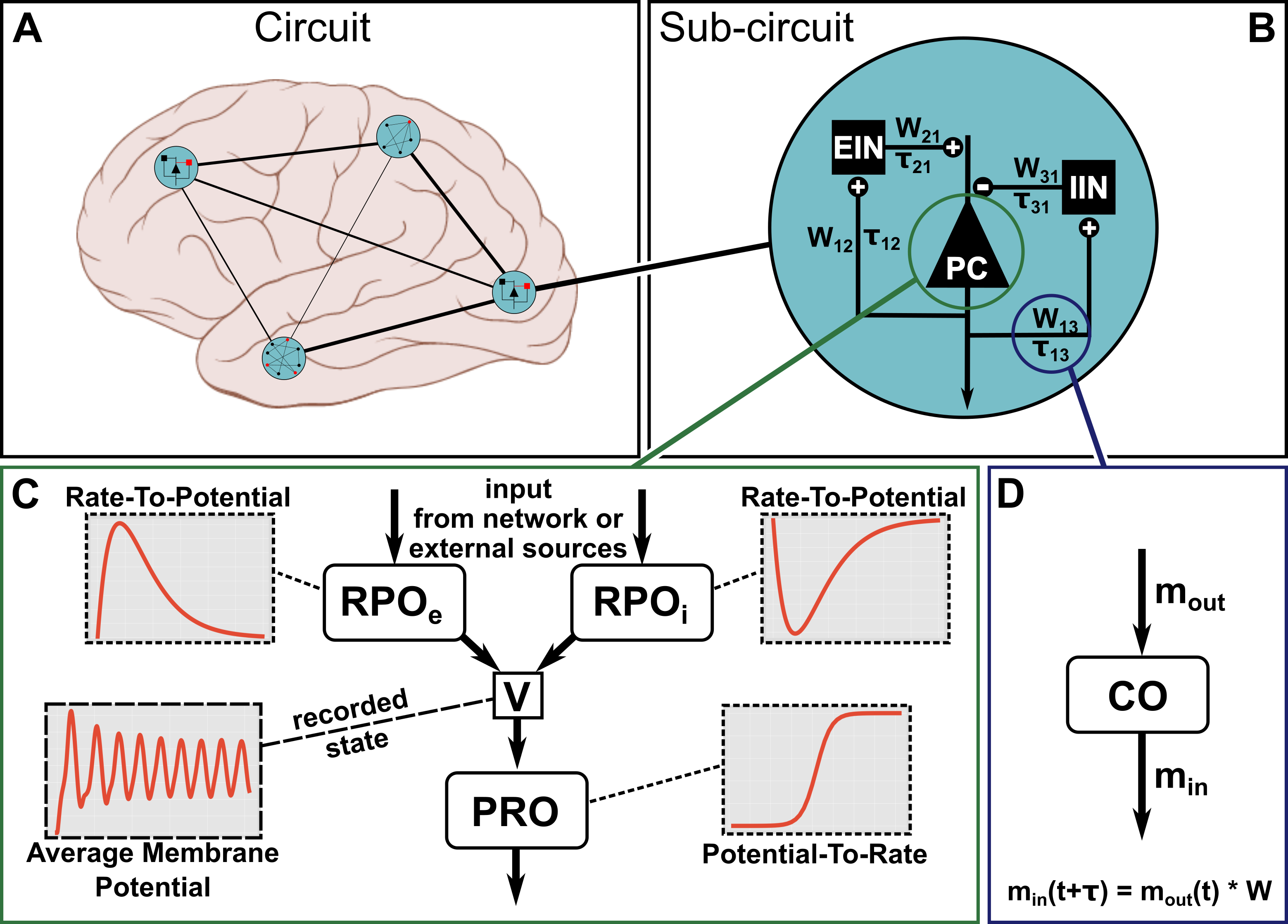
Figure 1
The structure of the full Jansen-Rit model is depicted in Figure 1 B. As visualized for the pyramidal cell population in Figure 1 C, the model can be decomposed into a number of generic mathematical operators that can be used to build the dynamic equations for the Jansen-Rit model. We will use this decomposition to introduce the different possibilities that exist in PyRates to compose the dynamic equations of a neural population from different operators. This will be done for all 3 nodes (i.e. populations), which will then be connected via edges to yield the full Jansen-Rit neural mass model. An alternative, single-operator way of the model definition will be introduced as well.
1.1.1. References
1.1.1.1. What is a YAML template file?
YAML is a data formatting specification that aims to be human-readable and machine-readable at the same time. It’s current installment can be thought of as an extension to the also popular JSON standard. PyRates uses YAML files to provide a simple and readable way to define network models without any Python code. Each yaml-based model definition will have to be placed inside a yaml file in order to be interpreted properly, i.e. the file needs to have the ending .yaml or .yml.
- Building models via YAML templates has several advantages:
the model definition requires a minimum of syntax
defining a model via YAML templates means setting up an independent, reusable model configuration file (the .yml file) of small size that can easily be stored, shared and read without any python knowledge
Each YAML template of a PyRates model structure can be reused as a base template for a new YAML template
However, if you know your way around Python and you prefer to set up your full workflow, from model definition to simulation, in Python, this is also possible in PyRates. Check out the Python-based model definition tutorial for this.
1.1.1.2. Part 1: Operator Templates
Operator templates are the way to define the governing mathematical equations of a model in PyRates. They are called operators, since the dynamic equations of neural models can often be decomposed into meaningful mathematical operators that can be re-used at multiple instances. By defining such mathematical operators as distinct operator templates in PyRates, these operators just have to be defined once and can then be used to define different parts of a model. We will now go through the 2 major operators that the Jansen-Rit model can be decomposed into.
1.1.2. Operator template for the PRO
Below you see a YAML template that defines the so-called potential-to-rate operator (PRO) used within each neural population of the Jansen-Rit model. As the name suggests, this operator transforms the average membrane potential within a population into an average firing rate. It is defined by the following instantaneous, sigmoidal transform:
In this equation, \(m_{out}\) and \(V\) represent the average firing rate and membrane potential, respectively, while \(m_{max}\), \(r\) and \(V_{thr}\) are constants defining the maximum firing rate, firing threshold variance and average firing threshold within the modeled population, respectively. A YAML template representation of this operator would look as follows:
PRO:
base: OperatorTemplate
equations: "m_out = m_max / (1. + exp(r*(V_thr - V)))"
variables:
m_out: output
V: input(0.0)
m_max: 5.0
r: 560.0
V_thr: 6e-3
As can be seen, this operator takes a membrane potential \(V\) as input, and returns a firing rate \(m_{out}\) as output. Its typical, sigmoidal shape can be seen in Figure 1 C.
1.1.3. Operator template structure:
As can be seen from this definition of an operator, each operator template requires 3 fields:
base, equations and variables.
baseindicates, which operator template to derive this template from
The default base for an operator template is the python class
OperatorTemplateIf you have other operator templates defined that are derived from
OperatorTemplate, you can use them as base as well. In this case, you will inherit all equations and variables of this operator template. You can add additional variables, or overwrite existing ones. Equations can only be added, but not overwritten.
equationscontains the defining equations of this operator
equations are defined as strings of characters
if the operator is defined by a single equation, you can just provide the string
If there is more than one equation, a list of string-based equations has to be provided. We will see an example of this later.
variablescontains the type and value definitions for each variable that appears in
equationseach variable has to be scalar
each variable definition starts with the name of the variable (i.e. the variable key)
possible values of a variable definition are:
variable-> for state variables which can change over time. The initial value can be specified in brackets, e.g.variable(0.1)input-> the variable will be provided with a value from a previous operator or external, user-defined inputoutput-> the value of this variable can be connected to another operatora scalar value, e.g.
m_max: 5.0-> indicates that this variable is a constant with value 5.0
be aware that there is a difference between the variable definition
a: 1and1.0, i.e. PyRates distinguishes between integers and float variables and requires matching variable types in mathematical operations.
1.1.4. Operator template for the RPO
The second important operator in a Jansen-Rit model is the rate-to-potential operator (RPO). It is conceptualized as convolution with an alpha kernel, which can be expressed as a second-order description of the synaptic response dynamics:
In these equations, \(V\) represents the average post-synaptic potential and \(H\) and \(\tau\) are the efficacy and the time-scale of the synapse, respectively. A PyRates YAML template for the RPO could look as shown below:
RPO_e:
base: OperatorTemplate
equations: ['d/dt * V = V_t',
'd/dt * V_t = H/tau * m_in - 2 * V_t/tau - V/tau^2']
variables:
V: output
I: variable
m_in: input
tau: 0.01
H: 0.00325
This is an example of an operator with multiple equations, which are provided as a list of strings. The operator takes
a firing rate \(m_{in}\) as input and returns a membrane potential \(V\) as an output. The unit response of
that operator is depicted in Figure 1 C. We use the sub-script e to denote that this operator defines the
synaptic response dynamic for an excitatory synapse. Since the PC population of the Jansen-Rit model expresses
inhibitory synapses as well (see Figure 1 B and C), we also need to define an inhibitory version of such a
synapse. This operator for this synapse will differ from the RPO_e operator only in the constant values for
\(H\) and \(\tau\), i.e. it will have a different strength and different decay rate. However, since the
governing equations will be equal, we can use the RPO_e operator as the base template:
RPO_i:
base: RPO_e
variables:
tau: 0.02
H: -0.022
Note, that we only re-defined the constants that needed to be changed, whereas everything else will be inherited from
RPO_e.
1.1.4.1. Part 2: Node Templates
Node templates are what is used in PyRates to define the dynamic equations of a network node (e.g. a neural population) via a hierarchy of operators, such as the ones defined above. Using the two operator templates defined above (PRO and RPO), each population of the Jansen-Rit model can be defined. As shown in Figure 1 B, there exist 3 of those: pyramidal cells (PCs), excitatory interneurons (EINs) and inhibitory interneurons (IINs). We will now define separate node templates for each population.
1.1.5. Node template for the EIN population
As can be seen in in Figure 1 B, the EIN population receives its only (excitatory) input from the PC population.
To model the dynamic changes in the membrane potential that are caused by the firing rate input from the PC
population, the RPO_e operator is used. Furthermore, the EIN population projects back to the PC population
via an excitatory synapse. To receive the average firing rate of the EIN population that is required to implement this
projection, the PRO operator has to be applied to the output of the RPO operator. This provides the operator hierarchy
that governs the role of the EIN population in the Jansen-Rit model. A node template of this population would look as
follows:
EIN:
base: NodeTemplate
operators:
- RPO_e
- PRO
1.1.6. Node template structure
In comparison to operator templates, node templates only require the definition of 2 fields:
base and operators.
basedefines which node template to derive this specific node template from
The default base for a node template is the python class
NodeTemplateIf you have other node templates defined that are derived from
NodeTemplate, you can use them as base as well. In this case, you will inherit all operators of this template. You can add additional operators, but not overwrite existing ones.
operatorscontains a list of names of previously defined operators
each list entry starts with a
-on a new lineoperator hierarchies are automatically derived from the
inputandoutputvariables of each operatorthus, the sequence in which the operators are placed inside the node template does not matter
however, circular dependencies between the operator inputs and outputs should be prevented (PyRates throws an error if such circularities are detected)
thus, an output variable on one operator, that should connect to the input variable of another operator, needs to have the same name as this input variable
1.1.7. Node template for the IIN population
As can be seen in Figure 1 B, the IIN population shows an identical connectivity to the PC population as the EIN population. Thus, it expresses an identical operator structure. The only difference between EIN and IIN population is how their projections back to the PC population affect the PC membrane potential, which is excitatory and inhibitory, respectively. Hence, we will define the IIN population template as follows:
EIN:
base: NodeTemplate
operators:
- RPO_e
- PRO
1.1.8. Node template for the PC population
Now, the center piece of the Jansen-Rit model is the PC population, which receives input from both the EIN and the IIN
population. Their synapses have opposing influences on its membrane potential (note the different signs of the
synaptic efficacies \(H\) used for the RPO_e and RPO_i operator definitions) and need to be
implemented via two separate operators that govern the synaptic dynamics of the EIN to PC and IIN to PC projections.
Otherwise, the structure of the PC population is the same as that of the EIN and IIN population:
PC:
base: NodeTemplate
operators:
- RPO_e
- RPO_i
- PRO
Since both the RPO_e and the RPO_i operator express an output variable \(V\) and the PRO
operator requires \(V\) as an input, PyRates will detect that there are multiple outputs mapping to a single
input variable. In such a case, a sum will be calculated over all output variables first, which is then provided as
input variable to the respective operator. In this specific example, the input \(m_{in}\) of the PRO operator on
the PC population will be calculated as \(m_{in} = V_e + V_i\) where \(V_e\) and \(V_i\) refer to the
output variables of the RPO_e and RPO_i operators, respectively.
1.1.8.1. Part 3: Edge Templates
Edge templates allow to define the dynamic equations for projections between nodes. For example, they could be used to
model axonal delay distributions via a convolution with a delay distribution function. For this purpose, PyRates
provides the EdgeTemplate base template. It follows exactly the same structure as a
NodeTemplate, i.e. it is defined via a base and a collection of operators. Since the Jansen-Rit
model uses very simple, linear projection operations (see the coupling operator CO in Figure 1 D), no edge
templates are required for this model. A detailed tutorial for how to implement different forms of edge operations
such as delays, convolutions etc., will be provided by the edge definitions example in this gallery.
1.1.8.2. Part 4: Circuit Templates
A circuit template is what is used in PyRates to combine a set of nodes and edges to a full network model.
1.1.9. A circuit template for the Jansen-Rit model
In the case of the Jansen-Rit model, this translates to connecting the PC, EIN and IIN populations via simple, linear
edges that can be set up within the CircuitTemplate as follows:
JRC:
base: CircuitTemplate
nodes:
EIN: EIN
IIN: IIN
PC: PC
edges:
- [PC/PRO/m_out, IIN/RPO_e/m_in, null, {weight: 33.75}]
- [PC/PRO/m_out, EIN/RPO_e/m_in, null, {weight: 135.}]
- [EIN/PRO/m_out, PC/RPO_e/m_in, null, {weight: 108.}]
- [IIN/PRO/m_out, PC/RPO_i/m_in, null, {weight: 33.75}]
1.1.10. Circuit template structure
A circuit template requires the definition of 3 fields: base, nodes and edges.
basedefines which circuit template to derive this specific circuit template from
The default base for a circuit template is the python class
CircuitTemplateIf you have other circuit templates defined that are derived from
CircuitTemplate, you can use them as base as well. In this case, you will inherit all nodes and edges of this template. You can add additional nodes and edges and overwrite existing nodes, but not edges.
nodeslists all nodes that this circuit is composed of
each node definition starts in a new line, with the name that is given to the node within the circuit, followed by the name of the node template that should be used (
name_within_circuit: name_of_node_template)
edgesif no edges exist in circuit, this field can be skipped
else, edges are defined by lists with four entries:
The source variable (PC/PRO/m_out refers to variable m_out in operator PRO of node PC)
The target variable
An edge template with additional operators (here, null means that no particular edge template is used).
A dictionary of variables and values that are specific to this edge.
more complex syntax can be used within the four entries to define more complex edges. A tutorial on how to use these will be provided by the edge definitions example in this gallery.
This concludes the YAML-based definition of the Jansen-Rit model in PyRates. To learn how to use this model definition to perform numerical simulations, check out the other examples in this gallery.